Numpy & Scipy - 1.6 The Solution of Matrix Equation (General Matrices)
- Numpy & Scipy - 1.1 Notation of Matrix and Vector, Matrix Input and Output
- Numpy & Scipy - 1.2 Convenient Functions of Matrix
- Numpy & Scipy - 1.3 Basic Manipulation of Matrices (1)
- Numpy & Scipy - 1.4 Basic Manipulation of Matrices (2)
- Numpy & Scipy - 1.5 Basic Manipulation of Matrices (3)
- Numpy & Scipy - 1.6 The Solution of Matrix Equation (General Matrices)
- Numpy & Scipy - 1.7 The Solution of Band Matrix
- Numpy & Scipy - 1.8 The Solution of Toeplitz Matrix and Circulant Matrix And How to Solve AX=B
- Numpy & Scipy - 1.9 Calculate Eigenvector and Eigenvalue of Matrix
Finding the Determinant
1
from scipy import linalg
- Import
linalgfromscipy.
- Use
linalg.det(Matrix)to calculate and return the determinant. - The underlying algorithm uses LU Decomposition: $\quad A = LU$
- The determinant calculation is executed as follows:
- Since all diagonal terms of $L$ are 1, $det(L)=1$, so we only need to calculate $det(U)$.
- Therefore, for the General Case, methods like Cramer’s Rule learned in linear algebra are not used.
1
2
3
4
5
6
7
8
9
A1 = np.array([[1,5,0],[2,4,-1],[0,-2,0]])
det = linalg.det(A1)
print(det)
A2 = np.array([[1,-4,2],[-2,8,-9],[-1,7,0]])
det = linalg.det(A2)
print(det)
1
2
3
-2.0
15.0
Why Solving Ax=b is Better Than Finding the Inverse
When finding the solution by calculating $A^{-1}$
$A^{-1}$ calculation time complexity: $\sim n^3$
$A^{-1}b$ calculation time complexity: $\sim n^2$
When finding the solution by solving the equation $A\mathbf{x} = \mathbf{b}$
Gauss Elimination: $\sim n^3$
LU decomposition:
- Time complexity for $A = LU$: $\sim n^3$
- Time complexity for solving $LU\mathbf{x} = \mathbf{b}$: $\sim n^2$
- Although both seem similar in terms of time complexity, solving the equation $A\mathbf{x} = \mathbf{b}$ is preferred over using the inverse matrix for the following reasons.
Numerical Accuracy
Computers represent numbers using floating points, which have limited precision. Calculating with an inverse matrix introduces more approximations compared to solving the equation directly, leading to inaccurate results. Also, inverse matrices are often avoided due to backward error analysis reasons.
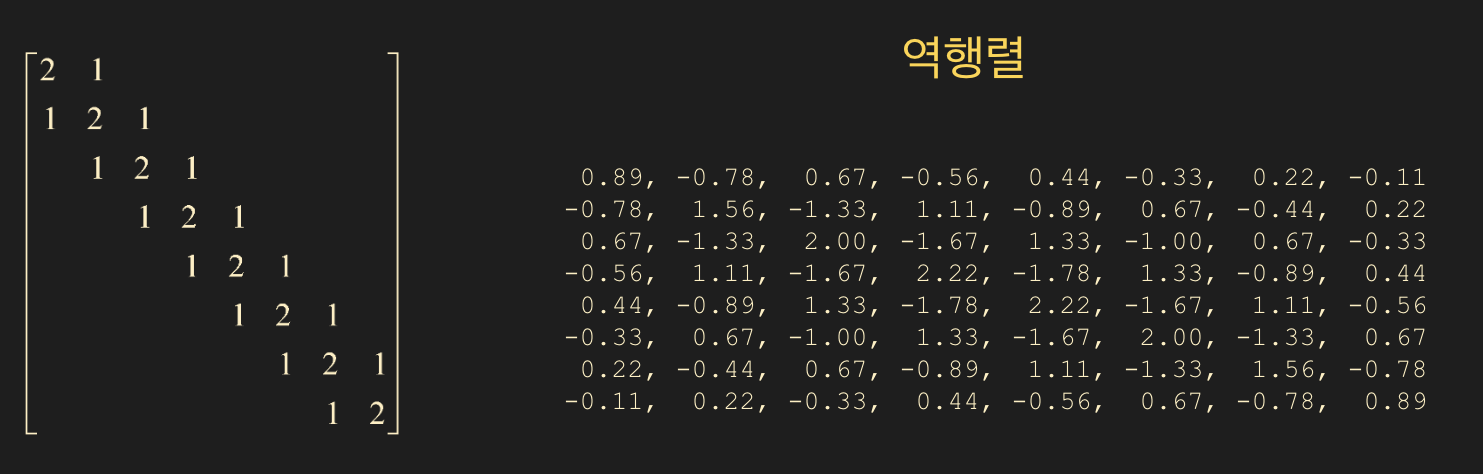
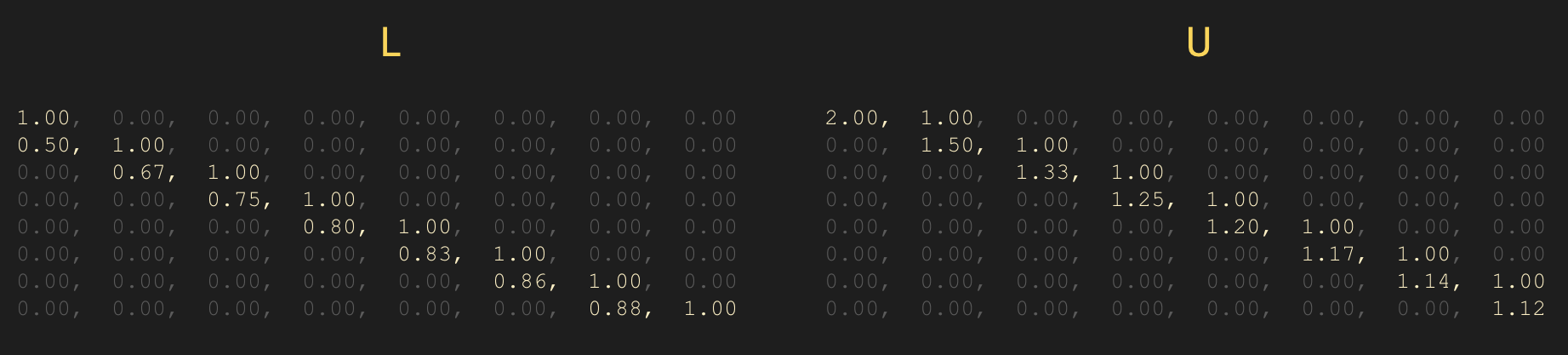
Comparison between Sparse Matrix Inverse and LU Decomposition
What happens to the inverse of a matrix if most of its elements are zero (sparse)?
- You might think the inverse of a sparse matrix would also be sparse, but reality isn’t that simple.
On the other hand, $L$ and $U$ often remain sparse.
- In summary, unless you specifically need the inverse matrix $A^{-1}$, do not calculate the inverse when solving $A\mathbf{x} = \mathbf{b}$!
Calculating the Inverse
- If your goal for finding the inverse is to solve $A\mathbf{x} = \mathbf{b}$, you should reconsider.
- Use
linalg.inv(matrix)to find the inverse matrix. If the input matrix is not invertible (i.e., it is a singular matrix), an error will occur.
- The basic algorithm uses LU decomposition and solves $\quad LUA^{-1} = I$, going through only the backward phase (back substitution).
1
2
3
4
A1 = np.array([[1,2,1],[2,1,3],[1,3,1]])
inv_a1 = linalg.inv(A1)
prt(inv_a1, fmt="%0.2f")
1
2
3
8.00, -1.00, -5.00
-1.00, 0.00, 1.00
-5.00, 1.00, 3.00
Solving the Matrix Equation Ax = b
Solving $A\mathbf{x} = \mathbf{b}$ is equivalent to finding $\mathbf{x}$.
- You can use the
x = linalg.solve(A, b, assume_a="gen")function. - The
assume_aparameter accepts four types of inputs.
gen
General solver. Used when the matrix type (symmetric, hermitian, etc.) is unknown.
Solves via LU decomposition: $\quad A = LU$
sym
Used for symmetric or complex symmetric matrices. Note that complex symmetric is not the same as Hermitian.
A symmetric matrix satisfies $\; A = A^T \;$.
Uses diagonal pivoting: $\quad A = LDL^T \mbox{(block diagonal)}$.
her
Used for Hermitian matrices: $\quad A = A^$ (conjugate transpose).
Solves via diagonal pivoting: $\quad A = LDL^$.
pos
Used for positive definite matrices. Implies the quadratic form $\mathbf{x}^TA\mathbf{x} > 0$. Uses Cholesky Decomposition: $\quad A = R^TR = LL^T$
- Be careful, as setting
assume_aincorrectly may not trigger an error but will return wrong results. Therefore,genis commonly used. - If you understand the matrix properties well, using the
assume_aoption is beneficial for computation speed.
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
b = np.ones((3,), dtype=np.float64)
# singular (not invertible)
A_sing = np.array([[1,3,4],[-4,2,-6],[-3,-2,-7]])
# gen (general)
A_gen = np.array([[0,1,2],[1,0,3],[4,-3,8]])
# sym (symmetric)
A_sym = np.array([[1,2,1],[2,1,3],[1,3,1]])
# sym complex (symmetric complex)
A_sym_c = np.array([[1, 2-1j, 1+2j],[2-1j, 1, 3],[1+2j, 3,1]])
# her (hermitian)
A_her = np.array([[1, 2+1j, 1-2j],[2-1j, 1, 3],[1+2j, 3,1]])
# pos (positive definite)
A_pos = np.array([[2, -1, 0],[-1,2,-1],[0,-1,2]])
- singular
- An error occurs because it is a singular matrix.
1
x = linalg.solve(A_sing, b)
1
numpy.linalg.LinAlgError: Matrix is singular.
- gen
- Added the last line to determine if $Ax = b ightarrow Ax - b = 0$.
1
2
3
4
5
x = linalg.solve(A_gen, b)
prt(x, fmt="%0.5e")
print()
prt(A_gen @ x - b , fmt="%0.5e")
1
2
3
1.00000e+00, 1.00000e+00, 0.00000e+00
0.00000e+00, 0.00000e+00, 0.00000e+00
- sym
- The difference between
symandgenlies in the underlying algorithm. - Keep in mind that the solution
xwe obtain is an approximation. Errors occur in values calculated using LU decomposition (gen) versus diagonal pivoting (sym).
1
2
3
4
5
6
7
8
9
10
x1 = linalg.solve(A_sym, b) # default is gen
x2 = linalg.solve(A_sym, b, assume_a="sym")
prt(x1, fmt="%0.5e")
print()
prt(A_sym @ x1 - b , fmt="%0.5e")
prt(x2, fmt="%0.5e")
print()
prt(A_sym @ x2 - b , fmt="%0.5e")
1
2
3
4
5
6
7
8
9
# gen
2.00000e+00, 0.00000e+00, -1.00000e+00
0.00000e+00, 0.00000e+00, 0.00000e+00
# sym
2.00000e+00, -2.77556e-17, -1.00000e+00
0.00000e+00, 0.00000e+00, 0.00000e+00
- her
1
2
3
4
5
x = linalg.solve(A_her, b, assume_a="her")
prt(x, fmt="%0.5e")
print()
prt(A_her @ x - b , fmt="%0.5e")
1
2
3
( 1.11111e-01+1.11111e-01j), ( 3.33333e-01-1.11111e-01j), ( 1.11111e-01+1.11022e-16j)
( 0.00000e+00+2.77556e-17j), (-4.44089e-16+1.94289e-16j), ( 2.22045e-16-1.66533e-16j)
- Here,
e-16means $10^{-16}$, which is close to 0.
- pos
1
2
3
4
5
6
7
8
9
10
x1 = linalg.solve(A_pos, b, assume_a="gen")
x2 = linalg.solve(A_pos, b, assume_a="pos")
prt(x1, fmt="%0.5e")
print()
prt(A_pos @ x1 - b , fmt="%0.5e")
prt(x2, fmt="%0.5e")
print()
prt(A_pos @ x2 - b , fmt="%0.5e")
1
2
3
4
5
6
7
8
9
# gen
1.50000e+00, 2.00000e+00, 1.50000e+00
0.00000e+00, 2.22045e-16, -4.44089e-16
# pos
1.50000e+00, 2.00000e+00, 1.50000e+00
-8.88178e-16, 1.55431e-15, -4.44089e-16
Triangular Matrix Solver
- This also solves $A\mathbf{x} = \mathbf{b}$, but only when matrix $A$ is a lower triangular or upper triangular matrix.
- Only the backward phase (back substitution) is required.
x = linalg.solve_triangular(A, b, lower=False)- Set the
lowerparameter toTruefor lower triangular andFalsefor upper triangular.
1
2
3
4
5
6
A = np.array([[1,0,0,0],[1,4,0,0],[5,0,1,0],[8,1,-2,2]], dtype=np.float64)
b = np.array([1,2,3,4],dtype=np.float64)
x = linalg.solve_triangular(A, b, lower=True)
prt(x, fmt="%0.5e")
1
1.00000e+00, 2.50000e-01, -2.00000e+00, -4.12500e+00
Is the Solution Accurate?
- The solution
xwe obtain when solving $A\mathbf{x} = \mathbf{b}$ is an approximation resulting from numerical calculations. Is $A\mathbf{x}$ similar enough to $\mathbf{b}$? This is equivalent to asking if $A\mathbf{x} - \mathbf{b}$ is close enough to 0.
- You can compare $A\mathbf{x} - \mathbf{b}$ with a zero matrix created via
np.zerosusing thenp.allclosefunction.
1
2
3
4
5
6
7
8
9
10
11
12
13
A = np.array([[2,-1,0],[-1,2,-1],[0,-1,2]], dtype=np.float64)
b = np.ones((3,), dtype=np.float64)
x = linalg.solve(A, b, assume_a="pos")
prt(x, fmt="%0.5e")
print()
prt(A@x-b, fmt="%0.5e")
zr = np.zeros((3,), dtype=np.float64)
bool_close = np.allclose(A@x-b, zr)
print(bool_close)
1
2
3
4
1.50000e+00, 2.00000e+00, 1.50000e+00
-8.88178e-16, 1.55431e-15, -4.44089e-16
True
Example 1.
$A = \begin{bmatrix} 1 & 1/2 & 1/3 & \dots & 1/9 & 1/10 \ 1/2 & 1/3 & 1/4 & \dots & 1/10 & 1/11 \ 1/3 & 1/4 & 1/5 & \dots & 1/11 & 1/12 \ \vdots & \ddots & \ddots & & & \ 1/9 & 1/10 & & & 1/17 & 1/18 \ 1/10 & 1/11 & \dots & \dots & 1/18 & 1/19 \end{bmatrix} $
Create a 10x10 Hilbert matrix using
linalg.hilbert(10)and store it inA.
Store the inverse ofAininv_A.
Calculatex1 = inv_A @ b.
Calculatex2usinglinalg.solve(gen).A@x1 - bwith 15 decimal places using custom output (floating).A@x2 - bwith 15 decimal places using custom output (floating).
CompareA@x1 - bandA@x2 - bwith a zero vector usingallclose.
- Solution
- The Hilbert matrix is numerically very difficult to handle. As its size increases, it becomes harder to solve accurately.
- Although we learned the theory, we can see that solving via inverse matrix is inferior in terms of accuracy.
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
20
21
22
23
24
25
26
A = linalg.hilbert(10)
b = np.ones((10,), dtype=np.float64)
# Hilbert Matrix
prt(A, fmt="%0.5f")
A_inv = linalg.inv(A)
x1 = A_inv @ b
print()
x2 = linalg.solve(A, b, assume_a="gen")
prt(A @ x1 - b, fmt="%0.15e")
print()
prt(A @ x2 - b, fmt="%0.15e")
print()
zr = np.zeros((10,), dtype=np.float64)
bool_close1 = np.allclose(A @ x1 - b, zr)
print(bool_close1)
bool_close2 = np.allclose(A @ x2 - b, zr)
print(bool_close2)
- Output of the Hilbert Matrix:
1 2 3 4 5 6 7 8 9 10
1.00000, 0.50000, 0.33333, 0.25000, 0.20000, 0.16667, 0.14286, 0.12500, 0.11111, 0.10000 0.50000, 0.33333, 0.25000, 0.20000, 0.16667, 0.14286, 0.12500, 0.11111, 0.10000, 0.09091 0.33333, 0.25000, 0.20000, 0.16667, 0.14286, 0.12500, 0.11111, 0.10000, 0.09091, 0.08333 0.25000, 0.20000, 0.16667, 0.14286, 0.12500, 0.11111, 0.10000, 0.09091, 0.08333, 0.07692 0.20000, 0.16667, 0.14286, 0.12500, 0.11111, 0.10000, 0.09091, 0.08333, 0.07692, 0.07143 0.16667, 0.14286, 0.12500, 0.11111, 0.10000, 0.09091, 0.08333, 0.07692, 0.07143, 0.06667 0.14286, 0.12500, 0.11111, 0.10000, 0.09091, 0.08333, 0.07692, 0.07143, 0.06667, 0.06250 0.12500, 0.11111, 0.10000, 0.09091, 0.08333, 0.07692, 0.07143, 0.06667, 0.06250, 0.05882 0.11111, 0.10000, 0.09091, 0.08333, 0.07692, 0.07143, 0.06667, 0.06250, 0.05882, 0.05556 0.10000, 0.09091, 0.08333, 0.07692, 0.07143, 0.06667, 0.06250, 0.05882, 0.05556, 0.05263
- Comparison of results from inverse matrix vs
np.solve(gen):
1
2
3
4
5
6
-5.384686879750245e-05, -4.824035864248177e-05, -4.365437722742005e-05, -3.984553430069759e-05, -3.663714443313815e-05, -3.390060580654719e-05, -3.154054138576612e-05, -2.948524204493541e-05, -2.767985240692550e-05, -2.608172012175114e-05
1.768403201651836e-10, 2.589735093039280e-10, 1.027888885118955e-11, -5.332401187274627e-13, -2.094682205466825e-10, -1.216354794664198e-10, 8.731149137020111e-11, 4.546962806273314e-11, 5.050826423769195e-11, -6.052347512053302e-11
False
True